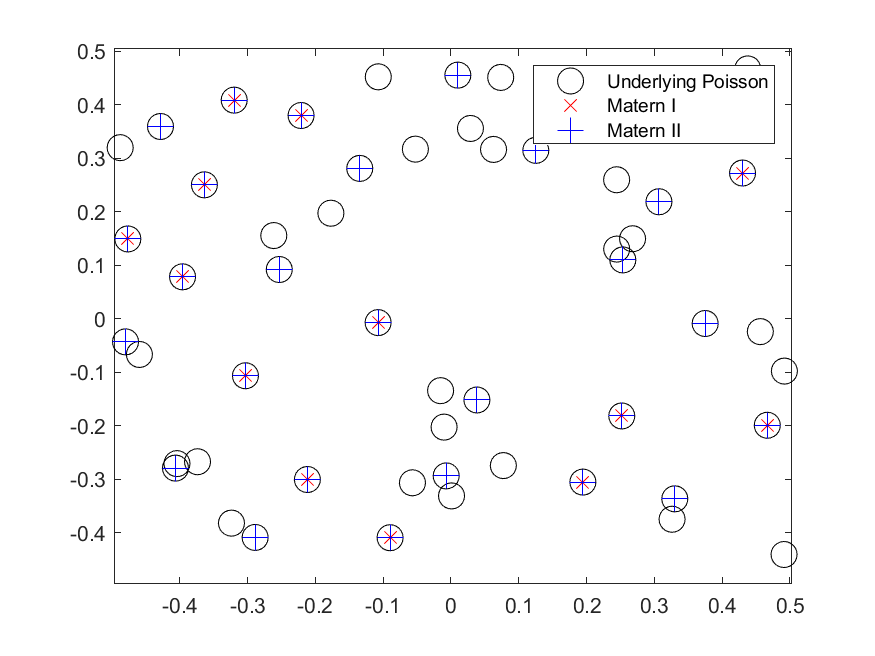
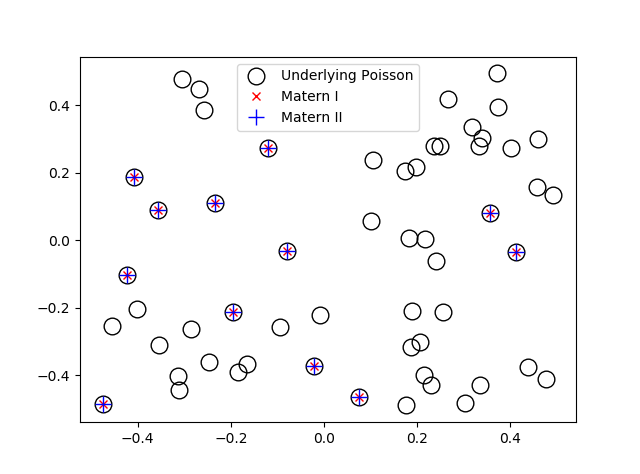
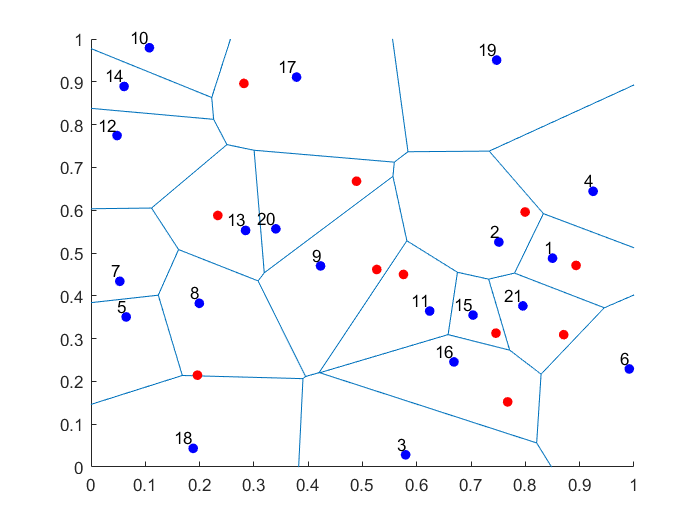
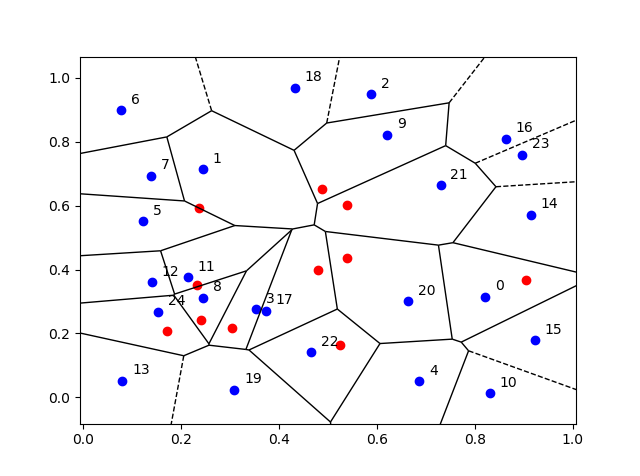
When sampling Poisson random variables, the method you use depends on the Poisson parameter, which coincides with its mean. The direct method, as I call it, is fine for small Poisson parameter values, but it becomes too slow as this parameter grows.
For large Poisson parameter values, researchers have proposed many other methods that are not slow. I coded up in Python and MATLAB (for now) two of these methods from the two respective papers:
- 1979, Atkinson, The computer generation of Poisson random variables;
- 1993, Hörmann, The transformed rejection method for generating Poisson random variable.
I already discussed these two methods in detail in a previous post, where you can find the code here.
The code I wrote is only suitable for large parameter values, where the meaning of large depends on the procedure being used, but it’s typically around 30. In other words, any code for generating Poisson variables should have an if-statement, using the direct method for small parameter values and another method, such as the ones above, for large parameter values.
Algorithms
Both algorithms are (acceptance-)rejection methods. I discuss this general method, first proposed by Neumann, in another post. In context of generating Poisson variables, these methods are, according to the book by Devroye (page 502), known for being relatively simple to implement and fast to execute.
Both algorithms require calculating the log of a factorial or, equivalently, a log of a gamma function, which is done by using approximations, such those of Stirling or Lanczos. I used the one by Stirling.
Algorithm PA by Atkinson (1979)
The Algorithm PA proposed by Atkinson, among other methods, is a rejection method that uses a logistic distribution as the envelope distribution. (Often such algorithms use a normal distribution as the envelope distribution.)
Algorithm PTRS by Hörmann (1993)
Hörmann refers to the Algorithm PTRS method as a transformed rejection method. It uses the inverse method, which I covered in a previous post, and a rejection method.
I have only seen one implementation of this algorithm. It’s written in C for the Python library NumPy; see the code here. You’ll notice my C code and that C code is very similar, modulus some logarithm identities.
Possible (small) error: I noticed that in that C code, on line 591, the implementation of step 2.0 of the PTRS Algorithm has a strictly less than condition, so it’s \(k <0\), whereas in the original paper (and hence my code), the condition is \(k\leq 0\). Perhaps this possible error is insignificant, as the procedure is for large-valued Poisson random variables, so the \(k=0\) scenario rarely happens.
Code
I used just a moment ratio to test the code. A proper test would involve a chi-squared test, for example, which is what I did when I wrote the same algorithms in Python and MATLAB; see this post here.
Warning: My C code is only for illustration purposes. If you can use an industrial library, use that.
Algorithm PA by Atkinson (1979)
/*
This code generates Poisson variates (or simulates Poisson variables).
using a method designed for large (>30) Poisson parameter values.
The generation method is Algorithm PA, a type of rejection method, from
the paper:
1979 - Atkinson - "The Computer Generation of Poisson Random Variables"
Author: H. Paul Keeler, 2020.
Website: hpaulkeeler.com
Repository: github.com/hpaulkeeler/posts
*/
static unsigned int funPoissonPA(double mu)
{
// precalculate some Poisson-parameter-dependent numbers
double c = 0.767 - 3.36 / mu;
double beta = pi / sqrt(3.0 * mu);
double alpha = beta * mu;
double k = log(c) - mu - log(beta);
double log_mu = log(mu);
// declare variables for the loop
double U, x, V, y, logfac_n, lhs, rhs;
unsigned int n;
unsigned int randPoisson = 0; // initialize the Poisson random variable (or variate)
bool booleContinue = true;
while (booleContinue)
{
U = funUniform(); // generate first uniform variable
x = (alpha - log((1.0 - U) / U)) / beta;
if (x < -.5)
{
continue;
}
else
{
V = funUniform(); // generate second uniform variable
n = floor(x + .5);
y = alpha - beta * x;
logfac_n = funLogFac(n);
// two sides of an inequality condition
lhs = y + log(V / (1.0 + exp(y)) / (1.0 + exp(y)));
rhs = k + n * log_mu - logfac_n; // NOTE: uses log factorial n
if (lhs <= rhs)
{
randPoisson = n;
booleContinue = false;
return randPoisson;
}
else
{
continue;
}
}
}
return randPoisson;
}Algorithm PTRS by Hörmann (1993)
/*
This code generates Poisson variates (or simulates Poisson variables).
using a method designed for large (>10) Poisson parameter values.
The generation method is Algorthm PTRS, a type of rejection method, from
the paper:
1993 - Hörmann - "The transformed rejection method for generating Poisson
random variables"
Author: H. Paul Keeler, 2020.
Website: hpaulkeeler.com
Repository: github.com/hpaulkeeler/posts
*/
/*WARNING:
This code uses rand(), the standard pseudo-random number generating function in C,
which is known for producing inadequately random numbers.
Replace the function rand() in the function funUniformSingle with another standard uniform number generator.
*/
static unsigned int funPoissonPTRS(double mu)
{
// precalculate some Poisson-parameter-dep}ent numbers
double b = 0.931 + 2.53 * sqrt(mu);
double a = -0.059 + 0.02483 * b;
double vr = 0.9277 - 3.6224 / (b - 2);
double one_over_alpha = 1.1239 + 1.1328 / (b - 3.4);
// declare variables for the loop
double U, V, us, log_mu, logfac_n, lhs, rhs;
unsigned int n;
unsigned int randPoisson = 0; // initialize the Poisson random variable (or variate)
bool booleContinue = true;
// Steps 1 to 3.1 in Algorithm PTRS
while (booleContinue)
{
// generate two uniform variables
U = funUniform();
V = funUniform();
U = U - 0.5;
us = 0.5 - fabs(U);
n = floor((2 * a / us + b) * U + mu + 0.43);
if ((us >= 0.07) && (V <= vr))
{
randPoisson = n;
return randPoisson;
}
if ((n <= 0) || ((us < 0.013) && (V > us)))
{
continue;
}
log_mu = log(mu);
logfac_n = funLogFac(n);
// two sides of an inequality condition
lhs = log(V * one_over_alpha / (a / us / us + b));
rhs = -mu + n * log_mu - logfac_n; // NOTE: uses log factorial n
if (lhs <= rhs)
{
randPoisson = n;
return randPoisson;
}
else
{
continue;
}
}
return randPoisson;
}Further reading
Algorithm PA was proposed in the paper:
- 1979, Atkinson, The computer generation of Poisson random variables.
This algorithm is also given in the book Monte Carlo Statistical Methods by Casella and Robert; see Algorithm A.6.
Algorithm PTRS was proposed in the paper:
- 1993, Hörmann, The transformed rejection method for generating Poisson random variable.